Discovering my gut microbiome
We have trillions of microbes in our gut, on our skin, and in virtually every nook and crevice of our body. In many ways, we are more microbial than we are human, and recent technologies have allowed us to characterize the thousands of different species that call our bodies home. By better understanding our microbial symbionts, we can better understand our health.
I’ve sequenced the fecal microbiome of dozens of individuals through my graduate research at the University of Illinois. Unfortunately, using your own specimens in the research lab is frowned upon, and living on a graduate stipend means I only have a limited budget for functional lab testing. So, three years went by, and I still didn’t know what was in my own gut!
A few months ago, I decided enough was enough. So, in January, I sent my sample to uBiome, a commercial stool microbiome testing company. Using the same state-of-the-art sequencing tools I use in the research lab, uBiome is helping people around the world learn about their microbiome.
Sample collection was quick and easy, and within weeks, I had my reports! Not only can you explore your microbiome at five relevant taxonomic levels (phylum, class, order, family, and genus), but you can also see the predicted functional capacity of your microbiome. You can even download the raw taxonomy data and get to the species level for some microbes.
Before I share my results, I should probably provide some context, since there are many things that can influence the composition of the fecal microbiome:
- I eat a low-carb, high-fat, paleo-type diet and regularly dip into ketosis, though I consumed a fair amount of carbs the day before this particular sample.
- I practice intermittent fasting and most days don’t eat until noon or 1 pm.
- I had pretty severe eczema prior to finding paleo and doing the autoimmune protocol. I have several food allergies/sensitivities.
- I had very bad asthma and upper respiratory infections as a kid and received many rounds of broad-spectrum antibiotics.
- I run or play soccer almost daily – aerobic exercise has been shown to have a significant impact on the microbiome.
- I still have some lingering SIBO/IBS, but I try not to restrict FODMAPs since many are prebiotics and help support a healthy colonic microbiome. I have been using antimicrobials but had taken a break from antimicrobial supplements at the time of sampling.
- I had also taken a break from any probiotic and prebiotic supplements at the time of sampling. I was occasionally consuming cooked and cooled white rice and potatoes (a source of resistant starch), and I always eat a wide variety of vegetables. Garlic and onion, which are potent prebiotics, are some of my favorite foods.
I also want to point out that there is still much to learn about the microbiome, so we don’t know enough to provide many actionable insights based solely on microbial abundances. Moreover, the fecal microbiome is not necessarily representative of the microbes that reside in the gut. Still, it’s certainly fun to peruse these data, and glean what we can from it.
Without further ado, here are the top 7 insights from finally getting my fecal microbiome sequenced:
1) My gut is slightly tilted towards Firmicutes
Firmicutes and Bacteroidetes are the two most abundant phyla in the human gut. While initial studies suggested that obesity was associated with a high Firmicutes to Bacteroidetes ratio, other studies have failed to find this association, and some more recent studies have even found the opposite association.1
2) I had LOTS of Akkermansia
Akkermansia is a genus with only one known species: Akkermansia muciniphila. This mucus-loving bacterium has been shown to be protective against weight gain and associated with reduced inflammation.2,3 I had 2.71 times more Akkermansia than most people. This may be at least in part due to the fact that I regularly practice intermittent fasting, as both fasting and caloric restriction have been shown to increase the abundance of A. muciniphila.4,5
3) My gut was fairly diverse
In general, higher gut diversity is associated with better health. My diversity score was 8.19 out of 10 (72nd percentile), which is pretty good, especially considering how many rounds of broad-spectrum antibiotics I had growing up. Interestingly, too much diversity can actually destabilize the gut ecosystem.
4) I had very few “probiotic” bacteria
I didn’t have any of the species typically associated with cheese, yogurt, raw milk, kefir, pickles, and sauerkraut, which is not entirely surprising. I used to have severe histamine intolerance, so my body doesn’t tolerate dairy, probiotics containing Lactobacillus and Bifidobacteria, or high amounts of fermented foods (other than kombucha) very well.
Interestingly, I had a few microbes that are associated with kimchi, though I don’t consume kimchi. I occasionally consume very small amounts of mixed fermented vegetables.
5) My top ten genera included several known butyrate-producers
Under the “My Bacteria” section, uBiome Explorer allows you to look at bacterial abundance at different taxonomic levels, with a visual pie chart. You can see my top ten genera below:
Probably the only criticism I have of uBiome is that I felt that their discussion of particular taxa was limited and in a few cases, not completely accurate. I’ll be talking about many of these microbes in detail in a future article series called “Meet your microbes”. For now, here’s a quick summary of my top 10 genera:
Bacteroides: This genus is known to be enriched in those who consume more animal protein and fat, and less carbohydrate.6 While Bacteroides species can use simple sugars when they are available, they prefer complex glycans from plants and dietary mucins.7
Faecalibacterium: This genus has only a single known species, Faecalibacterium prausnitzii, which breaks down complex carbohydrates and produces butyrate, which has widespread health benefits. This species is upregulated with exercise8 and may be protective against IBD. However, certain subspecies of F. prausnitzii may not be as health-promoting. One study found that the microbiome of atopic dermatitis patients was enriched subspecies of F. prausnitzii that may be less capable of producing butyrate.9 Since the uBiome sequencing depth only gets to the genus level, I don’t know which subspecies of Faecalibacterium are present in my gut.
Roseburia: digest complex carbohydrates and produce butyrate. Increased abundance of Roseburia is associated with improved glucose sensitivity and weight loss in mice.10 They are typically depleted in patients with IBS.
Blautia: this genus is known for its ability to combine hydrogen and carbon dioxide to form the short-chain fatty acid acetate. This may, in theory, be beneficial for reducing gas and bloating after the consumption of complex fibers that result in hydrogen and carbon dioxide production.11
Alistipes: a genus known to be increased with a diet high in animal protein and fat,12 and indicative of good gastrointestinal health. Alistipes is a butyrate-producer, and studies have found that high abundance of Alistipes is associated with a lower body weight.13
Subdoligranulum: not much is known about this genus, though it is supposedly reduced in people with IBS.11
Akkermansia: as discussed above, this mucus-loving bacterium has been shown to be protective against weight gain and associated with reduced inflammation.2,3
Parabacteroides: this genus is thought to be enriched in the presence of resistant starch or a high-fiber diet.14 It may protect against inflammation and is often reduced in IBS and IBD.15
Lachnospira: a genus in the Clostridium cluster XIVa composed of two species: Lachnospira multipara and Lachnospira pecinoschiza. These microbes are known to ferment pectin, a fiber found in many different fruits and vegetables, and produce acetate. This acetate may be converted to butyrate by other microbes via cross-feeding interactions.16
Pseudobutyrivibrio: this genus ferments a variety of carbohydrates to produce butyrate.
6) My microbiome is highly enriched for:
- Caffeine metabolism (7.67X higher than average): not surprising, as I was drinking a cup or two of home-roasted, fair trade Guatemalan coffee every day at the time of this sample!
- Steroid biosynthesis (6.98X higher than average): there are many types of steroids, so this is not too informative.
- Stilbenoid, diarylheptanoid, and gingerol biosynthesis (2.83X higher): these are secondary metabolites of plants. Gingerol is the active compound in fresh ginger.
- Flavonoid biosynthesis (2.15X higher): flavonoids are another type of secondary plant metabolites. Some have beneficial impacts on human health.
7) My microbiome is much less enriched for:
- Synthesis and degradation of ketone bodies (0.88X lower than average): this was surprising to me, given that I often dip in and out of ketosis, but I suppose it’s unlikely that many ketones produced in the liver end up in the gut lumen.
- D-Arginine and D-ornithine metabolism (0.51X lower than average): lactic acid bacteria (LAB) are major metabolizers of arginine and ornithine, so given that I had few LAB, this finding is not too surprising.
- Xylene and dioxin degradation (0.67X lower than average): xylenes and dioxins are ubiquitous environmental pollutants.
- Phagosome ability (0.01 X lower than average): the ability to engulf particles
Overall, I’m glad that I decided to send in a sample, and was impressed with the ease of sample collection and navigation of the uBiome Explorer interface. I look forward to the next iteration of commercial sequencing, which will likely include archaea, fungi, and even viruses.
That’s all for now! Over the next few months, I will be writing a series of articles called “Meet your microbes”, where I will talk about different microbial taxa, so if you choose to get your microbiome sequenced, keep the results handy, and subscribe to my weekly newsletter so you never miss a post.
Sources:
- Koliada, A. et al. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol. 17, (2017).
- Derrien, M., Belzer, C. & de Vos, W. M. Akkermansia muciniphila and its role in regulating host functions. Microb. Pathog. 106, 171–181 (2017).
- Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. 110, 9066–9071 (2013).
- Remely, M. et al. Increased gut microbiota diversity and abundance of Faecalibacterium prausnitzii and Akkermansia after fasting: a pilot study. Wien. Klin. Wochenschr. 127, 394–398 (2015).
- Sonoyama, K. et al. Response of Gut Microbiota to Fasting and Hibernation in Syrian Hamsters. Appl. Environ. Microbiol. 75, 6451–6456 (2009).
- David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
- Martens, E. C., Chiang, H. C. & Gordon, J. I. Mucosal Glycan Foraging Enhances Fitness and Transmission of a Saccharolytic Human Gut Bacterial Symbiont. Cell Host Microbe 4, 447–457 (2008).
- Allen, J. M. et al. Exercise Alters Gut Microbiota Composition and Function in Lean and Obese Humans. Med. Sci. Sports Exerc. (2017). doi:10.1249/MSS.0000000000001495
- Song, H., Yoo, Y., Hwang, J., Na, Y.-C. & Kim, H. S. Faecalibacterium prausnitzii subspecies–level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 137, 852–860 (2016).
- McGavigan, A. K. et al. TGR5 contributes to glucoregulatory improvements after vertical sleeve gastrectomy in mice. Gut 66, 226–234 (2017).
- Rajilić-Stojanović, M. et al. Intestinal microbiota and diet in IBS: causes, consequences, or epiphenomena? Am. J. Gastroenterol. 110, 278–287 (2015).
- De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. U. S. A. 107, 14691–14696 (2010).
- Verdam, F. J. et al. Human intestinal microbiota composition is associated with local and systemic inflammation in obesity. Obes. Silver Spring Md 21, E607-615 (2013).
- Martínez, I., Kim, J., Duffy, P. R., Schlegel, V. L. & Walter, J. Resistant Starches Types 2 and 4 Have Differential Effects on the Composition of the Fecal Microbiota in Human Subjects. PLoS ONE 5, (2010).
- Noor, S. O. et al. Ulcerative colitis and irritable bowel patients exhibit distinct abnormalities of the gut microbiota. BMC Gastroenterol. 10, 134 (2010).
- Esquivel-Elizondo, S., Ilhan, Z. E., Garcia-Peña, E. I. & Krajmalnik-Brown, R. Insights into Butyrate Production in a Controlled Fermentation System via Gene Predictions. mSystems 2, (2017).
Discovering my gut microbiome
We have trillions of microbes in our gut, on our skin, and in virtually every nook and crevice of our body. In many ways, we are more microbial than we are human, and recent technologies have allowed us to characterize the thousands of different species that call our bodies home. By better understanding our microbial symbionts, we can better understand our health.
I’ve sequenced the fecal microbiome of dozens of individuals through my graduate research at the University of Illinois. Unfortunately, using your own specimens in the research lab is frowned upon, and living on a graduate stipend means I only have a limited budget for functional lab testing. So, three years went by, and I still didn’t know what was in my own gut!
A few months ago, I decided enough was enough. So, in January, I sent my sample to uBiome, a commercial stool microbiome testing company. Using the same state-of-the-art sequencing tools I use in the research lab, uBiome is helping people around the world learn about their microbiome.
Sample collection was quick and easy, and within weeks, I had my reports! Not only can you explore your microbiome at five relevant taxonomic levels (phylum, class, order, family, and genus), but you can also see the predicted functional capacity of your microbiome. You can even download the raw taxonomy data and get to the species level for some microbes.
Before I share my results, I should probably provide some context, since there are many things that can influence the composition of the fecal microbiome:
- I eat a low-carb, high-fat, paleo-type diet and regularly dip into ketosis, though I consumed a fair amount of carbs the day before this particular sample.
- I practice intermittent fasting and most days don’t eat until noon or 1 pm.
- I had pretty severe eczema prior to finding paleo and doing the autoimmune protocol. I have several food allergies/sensitivities.
- I had very bad asthma and upper respiratory infections as a kid and received many rounds of broad-spectrum antibiotics.
- I run or play soccer almost daily – aerobic exercise has been shown to have a significant impact on the microbiome.
- I still have some lingering SIBO/IBS, but I try not to restrict FODMAPs since many are prebiotics and help support a healthy colonic microbiome. I have been using antimicrobials but had taken a break from antimicrobial supplements at the time of sampling.
- I had also taken a break from any probiotic and prebiotic supplements at the time of sampling. I was occasionally consuming cooked and cooled white rice and potatoes (a source of resistant starch), and I always eat a wide variety of vegetables. Garlic and onion, which are potent prebiotics, are some of my favorite foods.
I also want to point out that there is still much to learn about the microbiome, so we don’t know enough to provide many actionable insights based solely on microbial abundances. Moreover, the fecal microbiome is not necessarily representative of the microbes that reside in the gut. Still, it’s certainly fun to peruse these data, and glean what we can from it.
Without further ado, here are the top 7 insights from finally getting my fecal microbiome sequenced:
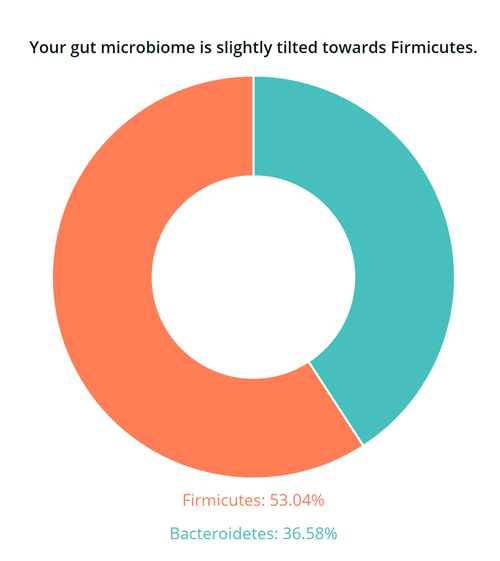
1) My gut is slightly tilted towards Firmicutes
Firmicutes and Bacteroidetes are the two most abundant phyla in the human gut. While initial studies suggested that obesity was associated with a high Firmicutes to Bacteroidetes ratio, other studies have failed to find this association, and some more recent studies have even found the opposite association.1
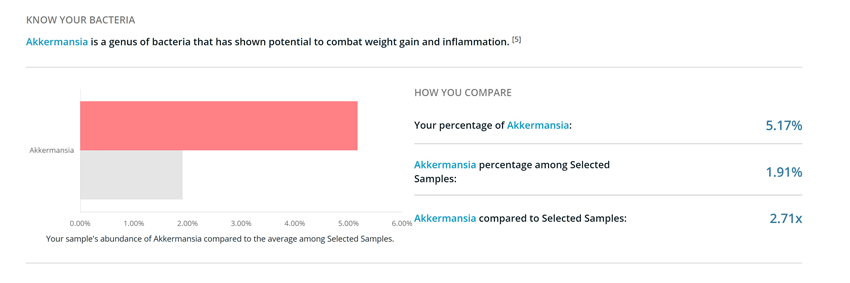
2) I had LOTS of Akkermansia
Akkermansia is a genus with only one known species: Akkermansia muciniphila. This mucus-loving bacterium has been shown to be protective against weight gain and associated with reduced inflammation.2,3 I had 2.71 times more Akkermansia than most people. This may be at least in part due to the fact that I regularly practice intermittent fasting, as both fasting and caloric restriction have been shown to increase the abundance of A. muciniphila.4,5
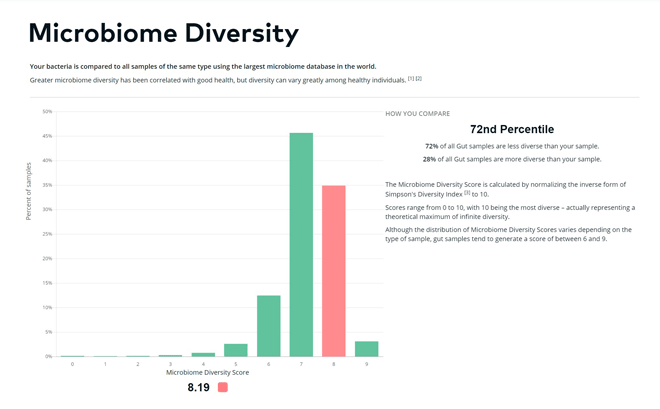
3) My gut was fairly diverse
In general, higher gut diversity is associated with better health. My diversity score was 8.19 out of 10 (72nd percentile), which is pretty good, especially considering how many rounds of broad-spectrum antibiotics I had growing up. Interestingly, too much diversity can actually destabilize the gut ecosystem.
4) I had very few “probiotic” bacteria
I didn’t have any of the species typically associated with cheese, yogurt, raw milk, kefir, pickles, and sauerkraut, which is not entirely surprising. I used to have severe histamine intolerance, so my body doesn’t tolerate dairy, probiotics containing Lactobacillus and Bifidobacteria, or high amounts of fermented foods (other than kombucha) very well.
Interestingly, I had a few microbes that are associated with kimchi, though I don’t consume kimchi. I occasionally consume very small amounts of mixed fermented vegetables.
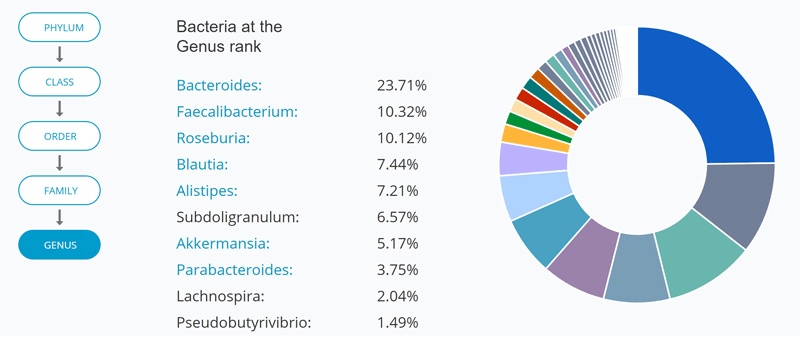
5) My top ten genera included several known butyrate-producers
Under the “My Bacteria” section, uBiome Explorer allows you to look at bacterial abundance at different taxonomic levels, with a visual pie chart. You can see my top ten genera below:
Probably the only criticism I have of uBiome is that I felt that their discussion of particular taxa was limited and in a few cases, not completely accurate. I’ll be talking about many of these microbes in detail in a future article series called “Meet your microbes”. For now, here’s a quick summary of my top 10 genera:
Bacteroides: This genus is known to be enriched in those who consume more animal protein and fat, and less carbohydrate.6 While Bacteroides species can use simple sugars when they are available, they prefer complex glycans from plants and dietary mucins.7
Faecalibacterium: This genus has only a single known species, Faecalibacterium prausnitzii, which breaks down complex carbohydrates and produces butyrate, which has widespread health benefits. This species is upregulated with exercise8 and may be protective against IBD. However, certain subspecies of F. prausnitzii may not be as health-promoting. One study found that the microbiome of atopic dermatitis patients was enriched subspecies of F. prausnitzii that may be less capable of producing butyrate.9 Since the uBiome sequencing depth only gets to the genus level, I don’t know which subspecies of Faecalibacterium are present in my gut.
Roseburia: digest complex carbohydrates and produce butyrate. Increased abundance of Roseburia is associated with improved glucose sensitivity and weight loss in mice.10 They are typically depleted in patients with IBS.
Blautia: this genus is known for its ability to combine hydrogen and carbon dioxide to form the short-chain fatty acid acetate. This may, in theory, be beneficial for reducing gas and bloating after the consumption of complex fibers that result in hydrogen and carbon dioxide production.11
Alistipes: a genus known to be increased with a diet high in animal protein and fat,12 and indicative of good gastrointestinal health. Alistipes is a butyrate-producer, and studies have found that high abundance of Alistipes is associated with a lower body weight.13
Subdoligranulum: not much is known about this genus, though it is supposedly reduced in people with IBS.11
Akkermansia: as discussed above, this mucus-loving bacterium has been shown to be protective against weight gain and associated with reduced inflammation.2,3
Parabacteroides: this genus is thought to be enriched in the presence of resistant starch or a high-fiber diet.14 It may protect against inflammation and is often reduced in IBS and IBD.15
Lachnospira: a genus in the Clostridium cluster XIVa composed of two species: Lachnospira multipara and Lachnospira pecinoschiza. These microbes are known to ferment pectin, a fiber found in many different fruits and vegetables, and produce acetate. This acetate may be converted to butyrate by other microbes via cross-feeding interactions.16
Pseudobutyrivibrio: this genus ferments a variety of carbohydrates to produce butyrate.
6) My microbiome is highly enriched for:
- Caffeine metabolism (7.67X higher than average): not surprising, as I was drinking a cup or two of home-roasted, fair trade Guatemalan coffee every day at the time of this sample!
- Steroid biosynthesis (6.98X higher than average): there are many types of steroids, so this is not too informative.
- Stilbenoid, diarylheptanoid, and gingerol biosynthesis (2.83X higher): these are secondary metabolites of plants. Gingerol is the active compound in fresh ginger.
- Flavonoid biosynthesis (2.15X higher): flavonoids are another type of secondary plant metabolites. Some have beneficial impacts on human health.
7) My microbiome is much less enriched for:
- Synthesis and degradation of ketone bodies (0.88X lower than average): this was surprising to me, given that I often dip in and out of ketosis, but I suppose it’s unlikely that many ketones produced in the liver end up in the gut lumen.
- D-Arginine and D-ornithine metabolism (0.51X lower than average): lactic acid bacteria (LAB) are major metabolizers of arginine and ornithine, so given that I had few LAB, this finding is not too surprising.
- Xylene and dioxin degradation (0.67X lower than average): xylenes and dioxins are ubiquitous environmental pollutants.
- Phagosome ability (0.01 X lower than average): the ability to engulf particles
Overall, I’m glad that I decided to send in a sample, and was impressed with the ease of sample collection and navigation of the uBiome Explorer interface. I look forward to the next iteration of commercial sequencing, which will likely include archaea, fungi, and even viruses.
That’s all for now! Over the next few months, I will be writing a series of articles called “Meet your microbes”, where I will talk about different microbial taxa, so if you choose to get your microbiome sequenced, keep the results handy, and subscribe to my weekly newsletter so you never miss a post.
Sources:
- Koliada, A. et al. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol. 17, (2017).
- Derrien, M., Belzer, C. & de Vos, W. M. Akkermansia muciniphila and its role in regulating host functions. Microb. Pathog. 106, 171–181 (2017).
- Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. 110, 9066–9071 (2013).
- Remely, M. et al. Increased gut microbiota diversity and abundance of Faecalibacterium prausnitzii and Akkermansia after fasting: a pilot study. Wien. Klin. Wochenschr. 127, 394–398 (2015).
- Sonoyama, K. et al. Response of Gut Microbiota to Fasting and Hibernation in Syrian Hamsters. Appl. Environ. Microbiol. 75, 6451–6456 (2009).
- David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
- Martens, E. C., Chiang, H. C. & Gordon, J. I. Mucosal Glycan Foraging Enhances Fitness and Transmission of a Saccharolytic Human Gut Bacterial Symbiont. Cell Host Microbe 4, 447–457 (2008).
- Allen, J. M. et al. Exercise Alters Gut Microbiota Composition and Function in Lean and Obese Humans. Med. Sci. Sports Exerc. (2017). doi:10.1249/MSS.0000000000001495
- Song, H., Yoo, Y., Hwang, J., Na, Y.-C. & Kim, H. S. Faecalibacterium prausnitzii subspecies–level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 137, 852–860 (2016).
- McGavigan, A. K. et al. TGR5 contributes to glucoregulatory improvements after vertical sleeve gastrectomy in mice. Gut 66, 226–234 (2017).
- Rajilić-Stojanović, M. et al. Intestinal microbiota and diet in IBS: causes, consequences, or epiphenomena? Am. J. Gastroenterol. 110, 278–287 (2015).
- De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. U. S. A. 107, 14691–14696 (2010).
- Verdam, F. J. et al. Human intestinal microbiota composition is associated with local and systemic inflammation in obesity. Obes. Silver Spring Md 21, E607-615 (2013).
- Martínez, I., Kim, J., Duffy, P. R., Schlegel, V. L. & Walter, J. Resistant Starches Types 2 and 4 Have Differential Effects on the Composition of the Fecal Microbiota in Human Subjects. PLoS ONE 5, (2010).
- Noor, S. O. et al. Ulcerative colitis and irritable bowel patients exhibit distinct abnormalities of the gut microbiota. BMC Gastroenterol. 10, 134 (2010).
- Esquivel-Elizondo, S., Ilhan, Z. E., Garcia-Peña, E. I. & Krajmalnik-Brown, R. Insights into Butyrate Production in a Controlled Fermentation System via Gene Predictions. mSystems 2, (2017).


Hello Lucy,
Good article thank you. I’ve been doing some research on Akkermansia muciniphila in an attempt to treat my MCAS and have mostly read that It has an anti-inflammation effect in the gut as you also state in your article, but I recently came across this research saying it causes inflammation. I’m interested in what you make of this?
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7831313/
Thanks
Hi Matt! We do sometimes see it elevated in states of inflammation, but this is most likely a consequence of the inflammation, rather than the cause of it. See this study, which indicates that Akkermansia expands in response to local gut injury, and helps to facilitate repair of the gut epithelium: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5076466/. My guess is that this is what is going on in the case of stroke as well – that Akkermansia expands in an attempt to repair damage to the gut. There may be some nuances though, which I’ll be covering in an article on Akkermansia in the near future!
The first paper you cite “Koliada, A. et al. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol. 17, (2017).” regarding your results for the F/B ratio actually confirms the commonly accepted axiom of BMI and F/B ratio are positively correlated. From their results section, “The obtained data indicate that obese persons in Ukraine adult population have a significantly higher level of Firmicutes and lower level of Bacteroidetes compared to normal-weight and lean adults.”
[…] are a poor indicator of gut microbial composition. This is why I don’t put too much stock in uBiome results, as much as I find them fascinating. I still believe comprehensive stool analysis is useful, as it […]